What is transcriptomics?
Transcriptomics is the study of all the RNA transcripts produced by an organism. Transcripts are produced from DNA and can give us an insight into gene expression levels in a cell [1]. RNA transcripts can provide the instructions for the cell to create a variety of proteins. The transcriptome is the collection of all the different RNAs in an organism, transcriptome databases can be useful for research on diseases such as OPA3. RNA-Seq and microarrays are molecular biology techniques performed in order to determine the transcriptome [2]. While RNA-Seq is the more expensive method, it has the ability to detect lower levels of transcripts and avoids technical errors experienced in microarrays.
OPA3 Human Transcriptome
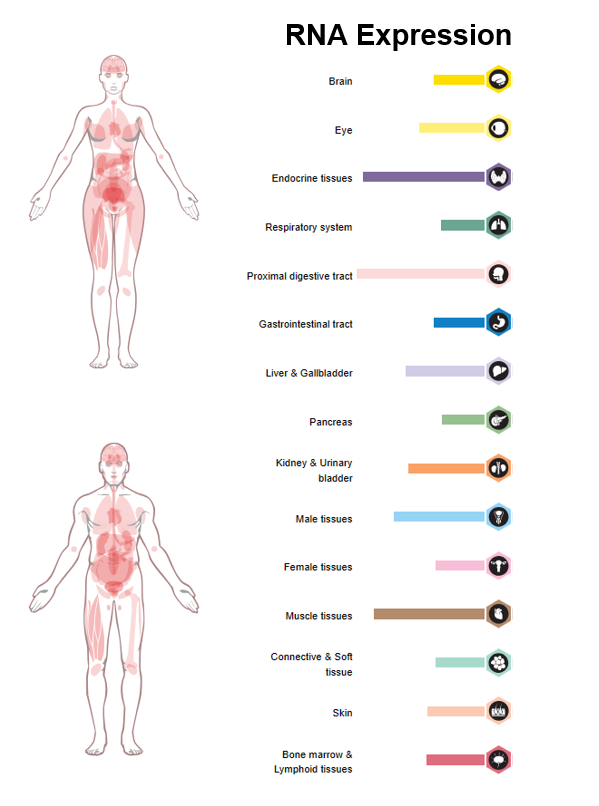
Figure 1. RNA Expression of OPA3 in females and males, taken from the Human Protein Atlas.
Discussion
In figure 1, above, the relative expression of OPA3 in the human body is visualized. The colorful bars represent the level at which RNA transcripts were found in the respective tissue. This figure provides evidence that OPA3 is very highly expressed in endocrine, proximal digestive tract, and muscles. Endocrine tissues are hormone producing tissues like the thymus, pituitary gland, thyroid and parathyroid glands.
From the figure we can see that OPA3 is expressed at a moderate level within the eye. It is likely that tissues in the eye are in some way more sensitive to changes in mitochondria morphology, although the mechanism by which this occurs is still unknown [3]. Studying the mitochondria's membrane dynamics in response to OPA3 mutations could help uncover the disease pathology for ADOA type 3.
From the figure we can see that OPA3 is expressed at a moderate level within the eye. It is likely that tissues in the eye are in some way more sensitive to changes in mitochondria morphology, although the mechanism by which this occurs is still unknown [3]. Studying the mitochondria's membrane dynamics in response to OPA3 mutations could help uncover the disease pathology for ADOA type 3.
This web page was produced as an assignment for Genetics 564, a capstone course at UW-Madison.
References
[1] Milward, E. A., Shahandeh, A., Heidari, M., Johnstone, D. M., Daneshi, N., & Hondermarck, H. (2016). Transcriptomics. In Encyclopedia of Cell Biology (pp. 160–165). Elsevier. https://doi.org/10.1016/B978-0-12-394447-4.40029-5
[2] Zhao, S., Fung-Leung, W.-P., Bittner, A., Ngo, K., & Liu, X. (2014). Comparison of RNA-Seq and Microarray in Transcriptome Profiling of Activated T Cells. PLoS ONE, 9(1), e78644. https://doi.org/10.1371/journal.pone.0078644
[3] Strachan, E. L., Mac White-Begg, D., Crean, J., Reynolds, A. L., Kennedy, B. N., & O’Sullivan, N. C. (2021). The Role of Mitochondria in Optic Atrophy With Autosomal Inheritance. Frontiers in Neuroscience, 15, 784987. https://doi.org/10.3389/fnins.2021.784987
Figure 1. The Human Protein Atlas - Accessed April 9th, 2024